PDX model details
| PDX ID |
410.51A |
| Host Strain(and Source) |
NSG
Source: Monash University |
| Host Strain Immune system Humanized |
NO |
| Host Type |
Testosterone supplemented |
| Graft Site |
Subcutaneous |
Current Generation
(* indicates number of generations grown in Castrate host) |
11 |
| Average PDX Generation Time (days +/- SEM) |
60 ± 7 |
| Tumour preparation |
Tumor solid |
| Tumour Characterization Technology |
Histology, IHC, STR profiling, targeted DNA sequencing, RNAseq |
| Tumour confirmed not to be of Mouse/EBV origin |
Yes; negative for CD45 and/or positive for human CK8/18 and/or human mitochondria by IHC |
| Passage QA performed |
Routine QA every 2-3 passages |
| Associated meta data |
| PDX model availability |
Yes (fixed, frozen) |
| Governance restriction for distribution |
Available to investigators with appropriate ethics approvals, approved EOI from MURAL committee, and material transfer agreements |
| Pubmed ID |
34413304 |
| |
|
| Markers |
410.51A |
| AR |
Y |
| PSA |
Y |
| PSMA |
Y |
| NE |
N |
| ERG |
N |
This heatmap displays the call type of curated CNVs in the presented samples.
This heatmap displays the call type of CNVs in commonly occurred samples.
This table displays curated CNVs
|
PDX ID
|
Gene Symbol
|
CNV Log2
|
CNV Copy
|
CNV Call
|
Experiments name
|
Platform
|
Reference genome
|
| 410.51A |
AKT3 |
1.22079 |
4.66149 |
gain
|
MURAL Prostate Cancer PDX collection |
Targeted GARVAN Panel |
hg19 |
| 410.51A |
BRAF |
0.694282 |
3.23616 |
gain
|
MURAL Prostate Cancer PDX collection |
Targeted GARVAN Panel |
hg19 |
| 410.51A |
CHD7 |
0.929286 |
3.80867 |
gain
|
MURAL Prostate Cancer PDX collection |
Targeted GARVAN Panel |
hg19 |
| 410.51A |
MYC |
0.929286 |
3.80867 |
gain
|
MURAL Prostate Cancer PDX collection |
Targeted GARVAN Panel |
hg19 |
| 410.51A |
KMT2A |
0.958529 |
3.88665 |
gain
|
MURAL Prostate Cancer PDX collection |
Targeted GARVAN Panel |
hg19 |
| 410.51A |
GNAS |
0.548247 |
2.92462 |
gain
|
MURAL Prostate Cancer PDX collection |
Targeted GARVAN Panel |
hg19 |
| 410.51A |
RB1 |
-0.761378 |
1.17987 |
loss
|
MURAL Prostate Cancer PDX collection |
Targeted GARVAN Panel |
hg19 |
| 410.51A |
RB1 |
-1.39432 |
0.760848 |
loss
|
MURAL Prostate Cancer PDX collection |
Targeted GARVAN Panel |
hg19 |
Clinical Information
| Sample Number |
410.51A |
| Sample Site |
Right lung sub pleural tumour |
| Sample source |
Autopsy |
| Pathology Tumour Diagnosis |
None |
| Gleason Score |
None |
| Primary Gleason Score |
None |
| Secondary Gleason Score |
None |
| Tertiary Gleason Score |
None |
| ISUP Grade Group |
|
| Tumour Grade |
|
| D'Amico Risk Classification |
|
| Tumour Volume (in cc) |
0.0 |
| Treatment Prior to Specimen Collection |
ADT, docetaxel, cabazitaxel, abiraterone, enzalutamide |
| |
|
Patient Information
| Patient Number |
410 |
| Sex |
Male |
| Diagnosis |
Prostate Cancer |
| PSA at diagnosis (ng/mL) |
30 |
| Consent to share data |
|
| |
|
This heatmap displays the mutations of curated sequence variants.
Download
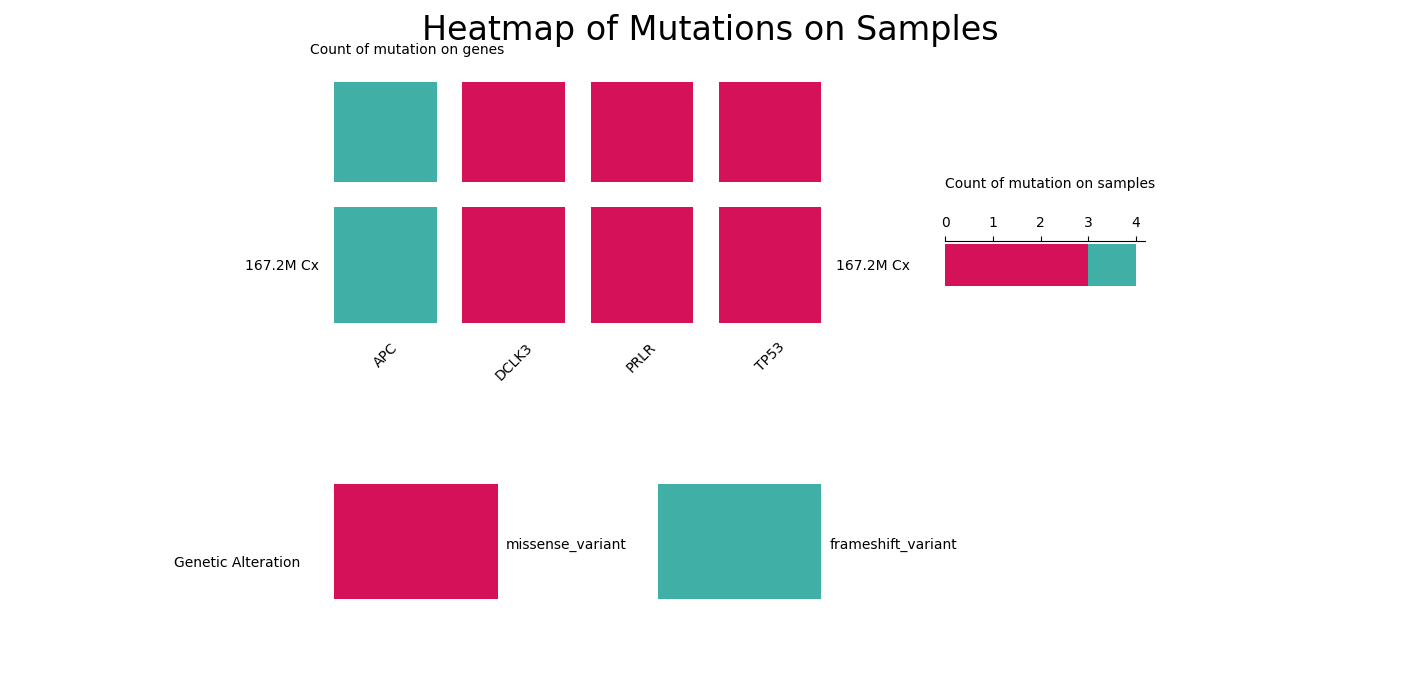
This table displays curated sequence variants
|
PDX ID
|
Gene Symbol
|
Depth
|
ALT_FREQ
|
Consequence
|
Exon
|
GnomAD_AF
|
CADD_PHRED
|
Clinvar_clnsig
|
Platform
|
Experiments Name
|
Reference
|
Library Type
|
Instrument Type
|
| 410.51A |
AKT3 |
485 |
0.81 |
missense_variant |
'3/14 |
. |
28.3 |
-
|
Targeted GARVAN Panel |
MURAL Prostate Cancer PDX collection |
hg19 |
paired |
Illumina HiSeq 2500 |
| 410.51A |
ZNF703 |
1030 |
0.51 |
missense_variant |
'1/2 |
0.0002802 |
28.6 |
-
|
Targeted GARVAN Panel |
MURAL Prostate Cancer PDX collection |
hg19 |
paired |
Illumina HiSeq 2500 |
| 410.51A |
GATA3 |
601 |
0.47 |
missense_variant |
'3/6 |
8.26E-06 |
26.1 |
-
|
Targeted GARVAN Panel |
MURAL Prostate Cancer PDX collection |
hg19 |
paired |
Illumina HiSeq 2500 |
| 410.51A |
CRLF2 |
462 |
0.31 |
missense_variant |
'6/6 |
. |
9.837 |
not_provided
|
Targeted GARVAN Panel |
MURAL Prostate Cancer PDX collection |
hg19 |
paired |
Illumina HiSeq 2500 |
No information in Drug dosing Table

